resources
We develop open-source tools for quantitative analysis of biological signals and single-cell dynamics.
Open Science & Methodological Assets
The following computational tools and experimental protocols were developed during the 2020–2025 Granada Lab cycle and remain available for the broader scientific community.
Code
Analysis code and computational tools developed in the course of our research are openly available on GitHub (Granada-Lab), including documentation and examples.
Datasets
Data underlying our publications are available in most publications and upon request. For access to specific datasets, annotated single-cell recordings, or analysis pipelines associated with a particular paper, please contact us directly.
Selected datasets will be deposited in public repositories as papers are published.
Movies and Talks
Talk at Single Cell Omics Germany
Invited talk by Adrián Granada at the Single Cell Omics Germany symposium. The talk covers quantitative single-cell approaches for studying cellular dynamics and treatment response in in-vitro cancer cell models.
Single Cell Circadian Oscillations
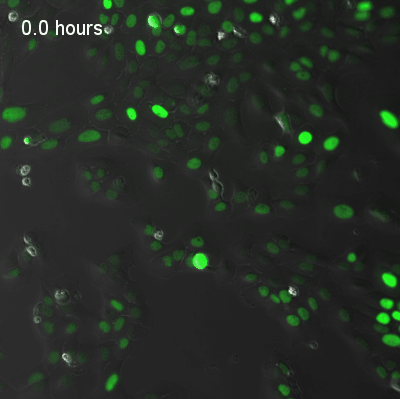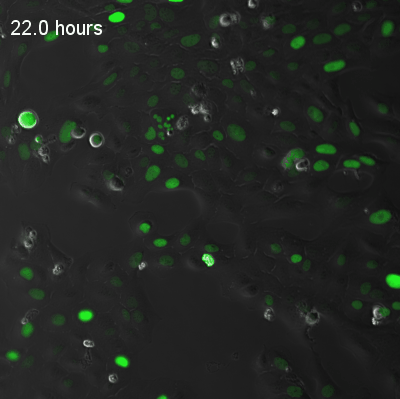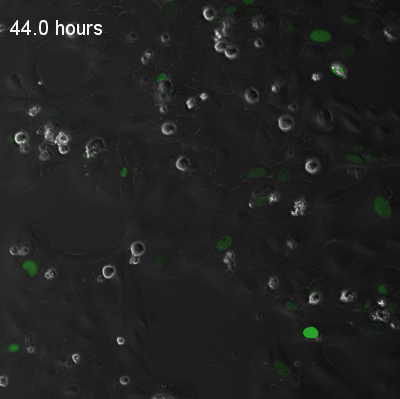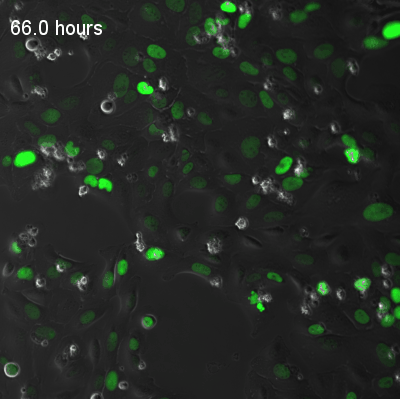
Live-cell fluorescence recordings of circadian clock reporter activity in individual U2OS cells. The two examples illustrate coupled and uncoupled dynamics at single-cell resolution:
Coupled cells — synchronised circadian oscillations across neighbouring cells.
Uncoupled cells — heterogeneous oscillatory dynamics with reduced intercellular coupling.
A step-by-step video guide for installing pyBOAT via the Anaconda Navigator. pyBOAT is an open-source Python package for time-frequency analysis of biological oscillations, including period estimation, amplitude extraction, and phase detection from noisy single-cell time series.
